Post-hoc testing in Kruskal-Wallis using R
Exercise 10.2.3.
Statistics for Ecologists (Edition 2) Exercise 10.2.3
These notes are about post hoc analysis when you use the non-parametric Kruskal-Wallis test. They supplement Chapter 10 in relation to using R to calculate differences between >2 non-parametric samples.
Post-hoc testing in the Kruskal-Wallis test using R
Introduction
The Kruskal-Wallis test is a non-parametric test for differences between more than two samples. It is essentially an analogue for a one-way anova. There is no “standard” method for carrying out post hoc analysis for KW tests. These notes show you how you can use a modified form of the U-test to carry out post hoc analysis.
When you carry out a Kruskal-Wallis test you are looking at the ranks of the data and comparing them. If the ranks are sufficiently different between samples you may be able to determine that these differences are statistically significant.
However, the main analysis only tells you about the overall situation, the result tells you that “there are differences”. Look at the following graph, which shows three samples.
A Kruskal-Wallis test of these data gives a significant result, H = 6.54 p < 0.05, but does not give any information about the pair by pair comparisons. Looking at the graph you might suppose that the Upper and Mid samples are perhaps not significantly different as their error bars (IQR) overlap considerably. The Lower sample appears perhaps to be different from the other two.
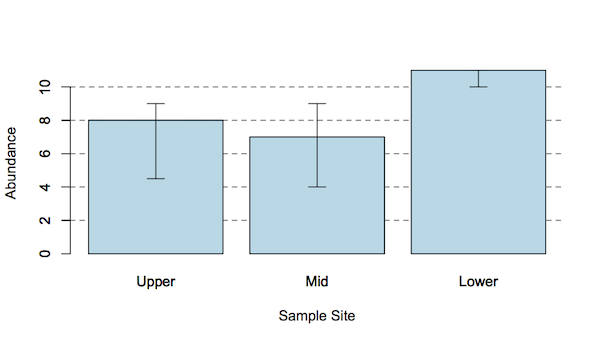
The way to determine these pairwise differences is with a post hoc test. You cannot simply carry out regular U-tests because you’ll need to carry out at least 3 and this “boosts” the chances of you getting a significant result by a factor of 3. You could simply adjust the p-values (e.g. using a Bonferroni correction), but this is generally too conservative.
These notes show you how to carry out a modified version of the U-test as a post hoc tool. The approach is analogous to the Tukey HSD test you’d use with parametric data.
A modified U-test as a post-hoc tool
With a bit of tinkering you can modify the formula for the U test to produce the following:
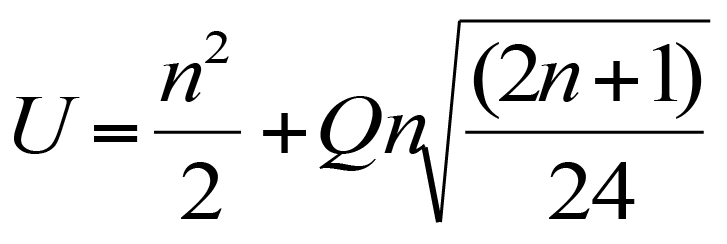
A critical value for U in a post hoc test for Kruskal-Wallis. Q is the Studentized range and n the harmonic mean of sample sizes.
In the formula n is the harmonic mean of the sample sizes being compared. and Q is the value of the Studentized Range for df = Inf, and number of groups equal to the original number of samples.
Critical values of Q, the Studentized range.
| Number of groups |
Significance | |
| 5% | 1% | |
| 2 | 2.772 | 3.643 |
| 3 | 3.314 | 4.120 |
| 4 | 3.633 | 4.403 |
| 5 | 3.858 | 4.603 |
| 6 | 4.030 | 4.757 |
The formula calculates a critical U-value for the pairwise comparison. You simply carry out a regular U-test, then use the largest U-value as a test statistic. If your value is equal or larger than the critical value from the formula, then the pairwise comparison is a statistically significant one.
Harmonic mean
The harmonic mean is easy to determine:
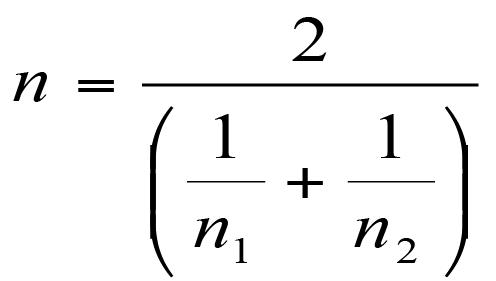
Calculating the harmonic mean of two values.
The harmonic mean is a way of overcoming differences in sample size. The more different the sample sizes the more unreliable this approach will be.
Calculate Q directly
If you re-arrange the formula you can calculate a value for Q:
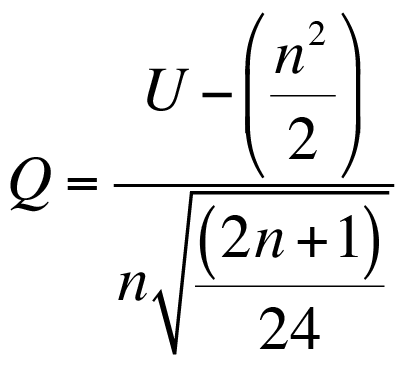
Calculate a value for Q (Studentized range) from the result of a U-test.
You can now use the result of a U-test (use the larger of the two calculated U-values) to work out a value for Q. Now you can compare your Q-value to the critical value.
The Studentized range is a distribution built into the basic R program. This gives you a way to compute an exact p-value for the pairwise comparison.
Custom R functions for Kruskal-Wallis post hoc
I’ve produced four custom functions for use with Kruskal-Wallis post hoc tests:
- mean() – Calculates the harmonic mean of two values.
- post() – Calculates the post hoc results for a pairwise comparison of two samples from a larger dataset.
- post() – Calculates the exact p-value for a post hoc given a U-value, number of groups and sample sizes.
- post() – Calculates a critical U-value for a post hoc given a confidence level (default 95%), number of groups and sample sizes.
These functions are contained in a single file: KW posthoc.R. If you source() the file you will set-up the functions and see a message giving some idea of what the functions do.
The h.mean() function
h.mean(n1, n2) |
|
| n1, n2 | Numerical values representing sample sizes of two samples. |
This function simply returns the harmonic mean of two numbers, i.e. the sample sizes of two samples.
h.mean(5, 7) [1] 5.833333
The function is called by the other post hoc functions (and is built-in) but it might be “handy” to have separately.
The KW.post() function
KW.post(x, y, data) |
|
| x, y | Numeric samples to compare. |
| data | The data object that contains the samples. It is assumed that the data is in sample format with multiple columns, each representing a separate sample. |
The function returns several results as a list:
- uval – The calculated U value for the pairwise comparison (the larger of the two calculated values).
- crit – The critical U value at 95%.
- value – The exact probability for the pairwise comparison.
The function also displays the results to the console, even if you assign the result to a named object.
hog3 Upper Mid Lower 1 3 4 11 2 4 3 12 3 5 7 9 4 9 9 10 5 8 11 11 6 10 NA NA 7 9 NA NA KW.post(Upper, Lower, data = hog3) Data: hog3 Pairwize comparison of: Upper and Lower U-value: 32.5 U-crit (95%): 31.06011 Post-hoc p-value: 0.02641968
If your data are in scientific recording layout, that is you have a response variable and a predictor variable, then you need a slightly different approach. You’ll have to work out a U-test result first, then run the KWp.post() and/or KWu.post() commands (see below).
The KWp.post() function
KWp.post(Uval, grp, n1, n2) |
|
| Uval | A calculated U-value for the pairwise comparison (the larger of the two calculated values). |
| grp | The number of groups (samples) in the original Kruskal-Wallis test. |
| n1, n2 | The sample sizes of the two groups being analysed. |
This function returns an exact p-value for a post hoc analysis. The value is returned immediately.
KWp.post(18, grp = 3, 7, 5)
Post-hoc p-value: 0.9851855
You can carry out a wilcox.test() on two samples to obtain a U-value. You need to know samples sizes because you need the larger of the two U values.
The wilcox.test() only gives one value so you must work out if the value you got was the largest. It so happens that:
n1 * n2 = U1 + U2
This means that you can work out the alternative U-value easily if you know sample sizes.
If your data are in recording layout, with a predictor and response, you’ll need to use the subset parameter to carry out the pairwise test:
hog2
count site
1 3 Upper
2 4 Upper
3 5 Upper
4 9 Upper
5 8 Upper
6 10 Upper
7 9 Upper
8 4 Mid
9 3 Mid
10 7 Mid
11 9 Mid
12 11 Mid
13 11 Lower
14 12 Lower
15 9 Lower
16 10 Lower
17 11 Lower
wilcox.test(count ~ site, data = hog2, subset = site %in% c("Upper", "Lower"))
Wilcoxon rank sum test with continuity correction
data: count by site
W = 32.5, p-value = 0.01732
alternative hypothesis: true location shift is not equal to 0
Check the sample sizes:
replications(count ~ site, data = hog2)
$site
site
Lower Mid Upper
5 5 7
Now you can see if the U-value you got was the largest:
5*7 – 32.5 [1] 2.5
Since it is the largest, you can use it in the KWp.post() function:
KWp.post(32.5, grp = 3, 5, 7) Post-hoc p-value: 0.02641968
Generally speaking the wilcox.test() will return the largest U-value if you use the response ~ predictor format for the command. If you run the command on separate samples the returned U-value will depend on the order you specify the samples.
The KWu.post() function
KWu.post(CI = 0.95, grp, n1, n2) |
|
| CI = 0.95 | The confidence interval, defaults to 0.95. This is essentially a significance level of p = 0.05. |
| grp | The number of groups (samples) in the original Kruskal-Wallis test. |
| n1, n2 | The sample sizes of the two groups being analysed. |
This function returns a U-value for a given confidence level. You supply the number of groups (samples) in the original Kruskal-Wallis test and the sizes of the two samples being compared. The result is a critical value, which means you can carry out the wilcox.test() and compare the resulting U-value to this critical value.
KWu.post(CI = c(0.95, 0.99), grp = 3, n1 = 5, n2 = 5)
Post-hoc critical U value: 23.71961 26.44729
In the example you see that you can set multiple confidence intervals. Here the critical U values for p = 0.05 and p = 0.01 are returned.
Comments are closed.